3'-5' Exonuclease
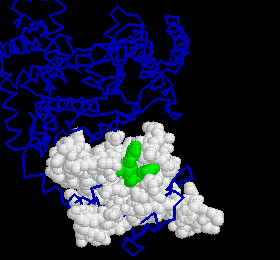

Fig. 1: DNA fragment in the 3'-5' active site. The DNA is represented in green, the active site in white and the blue backbone is the rest of the larger fragment.
The 3'-5' exonuclease activity of DNA Polymerase I is, like the polymerase active site, a part of the C-terminus segment, aka the Klenow Fragment. The entire enzyme is 928 amino acids long, with the Klenow fragment comprising those residues numbered 324 and higher. The 3'-5' site is situated between amino acids 324-500 (6).
The main function of the 3'-5' exonuclease site is proofreading newly synthesized DNA strands, assuring the accuracy of replication. The efficiency of this active site is such that the error rate of replication is virtually zero. As you can see in the figure below, the proximity of the 3'-5' exonuclease activity to the polymerase activity allows a quick transition between the editing and polymerizing modes.

Fig. 2: "P" indicates the location of the polymerase active site and "E" indicates the 3'-5' exonuclease site. The template strand enters through the top, is polymerized and, then, proofread by the exonuclease activity. Errors stall the polymerization, as the improperly bonded bases have to be excised from the newly synthesized, double-stranded DNA (10).
The 3'-5' exonuclease is postulated to catalyze the hydrolysis of a phosphosdiester bond with the aid of two metal ions. One of these divalent cations is proposed to help bind and orient both the single-stranded DNA substrate and an attacking water (3). The second cation plays the crucial role of acting as a Lewis acid to stabilize the negative charge that becomes resident on the leaving 3' oxygen. The second cation is essential for the proofreading reaction (3).
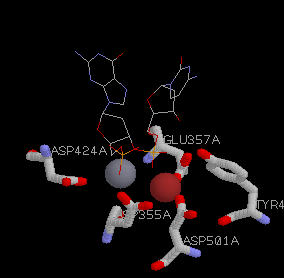
Fig. 3: Proposed 3'-5' mechanism with two metal ions. (a) The RasMol representation. The Mn2+ is pictured in gray and Zn2+ in red. Asp355 interacts with both of the metal ions, but not the DNA, directly .
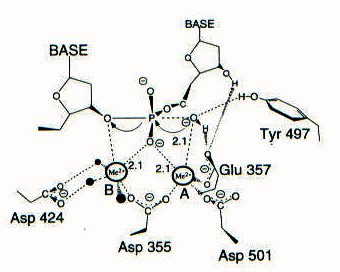
Fig. 4: An activated hydroxide attacks, having been oriented in the active site by Zn2+ (ion A). The pentacoordinated transitions state induces a negative charge to build on the leaving 3' atom, and this charge is stabilized by metal ion B (Mn2+) (3).
As you can see from the in Figs. 3 and 4, Zn2+, Glu357, and Tyr497 contribute to the orientation of the substrate and to the activation of the incoming nucleophile. The nucleophile attacks, generating a trigonal bipyramidal transition state, which is stabilized by both metals. The second metal ion also stabilizes the negative charge that comes to reside on the leaving 3' oxygen, thus facilitating its departure from the phosphate. This mechanism appears to be common to all proofreading domains of polymerases (2).